Moreover, quantification of the concentrations of C-reactive protein and prostate-specific antigen was illustrated by coupling the GIS with standard additions of purified protein standards. The utility of (18)O- labeled GIS was further illustrated by accurate relative quantification of 45 major human plasma proteins. coefficient of variance <10%) for analyte concentrations that were set at 100-fold higher or lower than those of the GIS based on the light ((16)O)/heavy ((18)O) peak area ratios. Reliable quantification was observed with high reproducibility (i.e. The accuracy, reproducibility, and linear dynamic range of quantification were further assessed based on known ratios of standard proteins spiked into the labeled mouse plasma reference. Our results showed that the percentage of heavy isotope ((18)O) incorporation applying an improved protocol was >99.5% for most peptides investigated. The (18)O- labeled proteome reference (or GIS) can be readily prepared and contains a heavy isotope ((18)O)- labeled internal standard for every possible tryptic peptide. Herein we present a proof of concept study using an (18)O- labeled proteome reference as global internal standards (GIS) for SRM-based relative quantification. Selected reaction monitoring (SRM)-MS is an emerging technology for high throughput targeted protein quantification and verification in biomarker discovery studies however, the cost associated with the application of stable isotope- labeled synthetic peptides as internal standards can be prohibitive for screening a large number of candidate proteins as often required in the preverification phase of discovery studies. 
Kim, Jong-Seo Fillmore, Thomas L Liu, Tao Robinson, Errol Hossain, Mahmud Champion, Boyd L Moore, Ronald J Camp, David G Smith, Richard D Qian, Wei-Jun PMID:26477253ġ8O- labeled proteome reference as global internal standards for targeted quantification by selected reaction monitoring-mass spectrometry. Sample preparation protocols including spin- labeling methods and preparation of membrane mimetic systems are also described. We focus on experimental approaches to quantify peptide-membrane binding, topology of bound peptides, and characterize peptide aggregation. In this chapter, we describe basic approaches for using SDSL EPR spectroscopy to study interactions between small peptides and biological membranes or membrane mimetic systems. While the method requires an introduction of a paramagnetic probe at a well-defined position in a peptide sequence, it has been shown to be minimally destructive to the peptide structure and energetics of the peptide-membrane interactions. The growth is driven by development of labeling strategies, as well as by considerable technical advances in the field, that are paralleled by an increased availability of EPR instrumentation. Site-directed spin labeling (SDSL) in combination with Electron Paramagnetic Resonance (EPR) spectroscopy is a well-established method that has recently grown in popularity as an experimental technique, with multiple applications in protein and peptide science. Peptide-membrane Interactions by Spin- labeling EPR 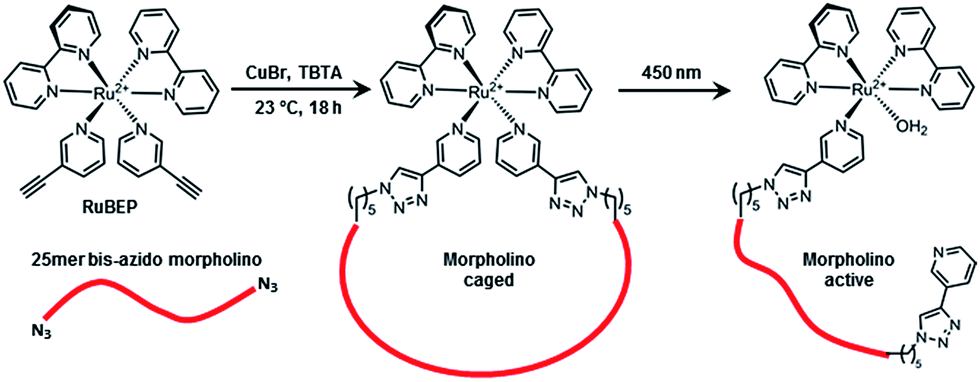
Our study suggests that the combinatorial labeling of peptides is a useful method to validate protein identifications for high confidence protein inference. Dual labeling has significantly reduced the false positive protein identifications in standard bovine six peptide digest. The improved representation has allowed us to accurately annotate the peptide sequences de novo. The dual modification resulted in improved fragment ion occurrence and intensity changes, and this resulted in the equivalent representation of b- and y-type fragment ions in an ion trap MS/MS spectrum. In this study, the influence of acetylation, guanidinylation, and their combination on peptide fragmentation was assessed initially on a lipase (LipA) from Bacillus subtilis followed by a bovine six protein mix digest. Particularly, chemical derivatizations of peptides were known to alter their fragmentation. Occurrence and intensity of these fragment ions in the MS/MS spectra are dependent on many factors such as amino acid composition, peptide basicity, activation mode, protease, etc. Identification of the protein from a set of experimental peptide spectral matches is usually referred as protein inference. Peptide mass spectra are obtained upon gas-phase fragmentation. Nalam, Madhusudhana RaoĪnnotation of peptide sequence from tandem mass spectra constitutes the central step of mass spectrometry-based proteomics. Kuchibhotla, Bhanuramanand Kola, Sankara Rao Medicherla, Jagannadham V. Combinatorial Labeling Method for Improving Peptide Fragmentation in Mass Spectrometry 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
